Experiment summary
Overview
This experiment is designed to find out stuff about memory.
Session Types
Each day, each mouse participated in one or more sessions. E.g. in the most common day, a mouse partakes in a single VR session.
Data
Data overview
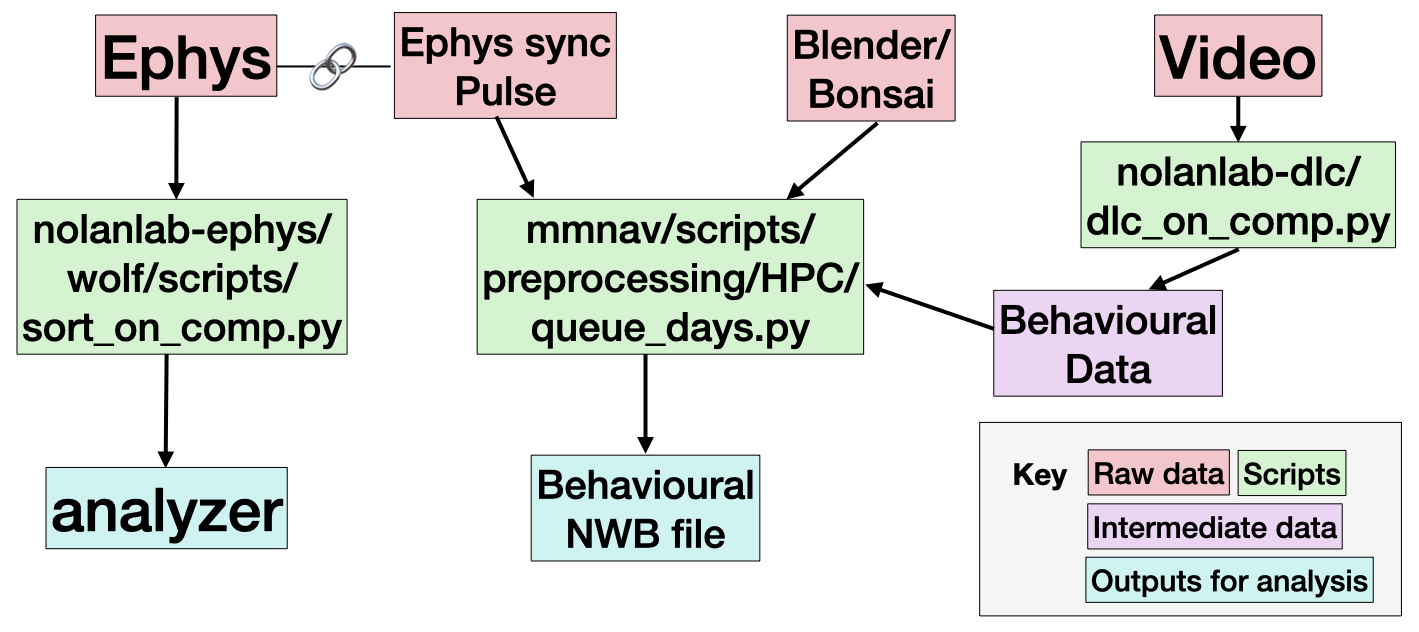
Our basic pipeline overview can be found below.
Every session contains an output from Bonsai or Blender, and a Video. These capture the animal behaviour data, as well as a light pulse signal used for synchronisation. Most sessions include ephys data, which capture neural behaviour.
We expect the most useful data to be the outputs for analysis (in teal), or the raw data (in red).
Data structure
The raw data is organised as follows:
raw/
session_folder/
M{mouse}_D{day}_datetime_{session_name}/
M{mouse}_D{day}_{session_name}_{datetime}.log # ??
M{mouse}_D{day}_{session_name}_{video_name}.avi # video of behaviour
M{mouse}_D{day}_{session_name}_{video_name}.csv # behavioural output from Bonsai/Blender
Record Node 109/ # Ephys dataThe processed data is organised as follows:
processed/
M{mouse}/
D{day}/
{session_name}/
sub-{mouse}_day-{day}_ses-{session_name}_srt-{sorter_protocol}_analyzer.zarr # a spikeinterface SortingAnalyzer
sub-{mouse}_day-{day}_ses-{session_name}_srt-{sorter_protocol}_{curation_protocol}.json # a file detailing the curation of the units (e.g. are they good units? Possible merges etc)
sub-{mouse}_day-{day}_ses-{session_name}_beh.nwb # an nwb file containing the behavioural outputsThe sorting output depends on a sorting_protocol, e.g. kilosort4A. The protocols are specified in github:chrishalcrow/nolanlab-ephys/src/nolanlab_ephys/si_protocols.py.
Reading the data
Behavioural data
The behavioural data output is designed to easy use with the pynapple package. You load it like so
import pynapple as nap
beh = nap.load_file('sub-04_day-08_ses-MMNAV2_beh.nwb')
print(beh['P']) # gives the position on the VR trackAnalyzer
The spike sorting output is stored in a SpikeInterface SortingAnalyzer. This can be loaded like so
import spikeinterface.full as si
analyzer = si.load_sorting_analyzer('sub-04_day-08_ses-MMNAV2_srt-kilosort4A_analyzer.zarr')We also save curation json files, which can be applied to the analyzer:
curation = si.load_curation('sub-04_day-08_ses-MMNAV2_srt-kilosort4A_curationA.json')
curated_analzyer = analyzer.apply_curation(curation)The analyzer can be easily convert to a pynapple object, to be easily used alongside the behavioural data:
spikes = si.to_pynapple_tsgroup(curated_analzyer)Note that the analyzer can easily be viewed using SpikeInterface-GUI. You need to install it, then you can run e.g.
sigui path/to/analyzer --curation --curation-file path/to/curation/fileThis allows you to check the automated curation we apply.
Raw ephys
You might want to look at the raw data directly. You can open it using spikeinterface
recording = si.read_openephys('M03_D09_2025-11-20_13-58-15_OF1')Running the pipeline
The basic pipeline contains three scripts. We’ll go through how to run these on the Edinburgh EDDIE compute cluster here.
Sorting
Install chrishalcrow/nolanlab-ephys following the instructions here: https://github.com/chrishalcrow/nolanlab-ephys .
For a sorting, you need to specify a mouse, a day, the sessions you want to sort and the sorting protocol to use. You also need to point to your raw data folder and your derivatives folder. To run a sorting, you would run a command such as the one below from the nolanlab-ephys directory:
uv run scripts/wolf/sort_on_eddie.py 4 8 MMNAV1 kilosort4A --data_folder /exports/eddie/scratch/chalcrow/wolf/data/ --deriv_folder /exports/eddie/scratch/chalcrow/wolf/derivativesThe exact same arguments can be used locally with a different script:
uv run scripts/wolf/sort_on_comp.py 4 8 MMNAV1 kilosort4A --data_folder /exports/eddie/scratch/chalcrow/wolf/data/ --deriv_folder /exports/eddie/scratch/chalcrow/wolf/derivativesDeepLabCut
Install chrishalcrow/nolanlab-dlc following the instructions here: https://github.com/chrishalcrow/nolanlab-dlc .
For a deeplabcut run, you need to stage the DLC model used to the EDDIE, then change its path in the config.yaml file. This is annoying. The models can be found in DATASTORE/ActiveProjects/Harry/deeplabcut/.
You need to specify a mouse, a day, a session and a bodypart. A typical command looks like:
uv run dlc_on_eddie.py 4 8 MMNAV1 tongue --data_folder /exports/eddie/scratch/chalcrow/wolf/data/ --deriv_folder /exports/eddie/scratch/chalcrow/wolf/derivativesDLC will be much more efficient if you specify a region of interest. These are stored in e.g. nolanlab-dlc/wolf_crops/tongue_crops_wolf.csv.
You can also run things locally using e.g.
uv run dlc_on_comp.py 4 8 MMNAV1 tongue --data_folder /exports/eddie/scratch/chalcrow/wolf/data/ --deriv_folder /exports/eddie/scratch/chalcrow/wolf/derivativesSync and preprocess
The syncing and preprocessing of data is done using https://github.com/wulfdewolf/mmnav . Install this and follow the instructions about environments.
You need to specify a mouse, a day and sorter protocol. The sessions are determined automatically. If the day has no ephys, don’t include a sorting protocol. A typical command looks like:
UV_PROJECT_ENVIRONMENT=$MMNAV_ENV uv run --no-sync --link-mode=copy --cache-dir $STORAGE/code/uv_cache/ scripts/preprocessing/HPC/queue_days.py --storage $STORAGE --sub 4 --day 15 --srt kilosort4A